Co-Analysis of AMP PDRD, GP2, & CRN Cloud Data
Verily Workbench is a cloud-based platform designed for collaborative biomedical research. It integrates data management, analysis tools, and governance controls within a unified interface.
Key features include:
- Workspaces
- Workspaces are where researchers and teams connect, collaborate, and organize all the elements of their research, including data, documentation, code, and analysis.
- Cloud apps
- Cloud apps are a configurable pool of cloud computing resources. They’re ideal for interactive analysis and data visualization, and can be finely tuned to suit analysis needs.
- Data Collections
- Data collections represent multimodal datasets that you can publish to Verily Workbench’s data catalog, so users can reference these data in workspaces.
- Access, browse, save, and share data
- Verily Workbench provides a variety of features to browse and interact with data in your workspace, making accessing, browsing, saving, and sharing your data easy.
What's on this page:
Pilot Program
Summary/Overview
To support your explorations, each program offers “Getting Started” workspaces within Workbench, providing sample notebooks and guidance tailored to their respective datasets. This companion document outlines the specific steps and resources necessary to use the AMP PDRD, GP2, and CRN Cloud datasets together.
Workspaces and Data Resources
The following workspaces and data catalogs are available to support your explorations. These resources provide program-specific examples, documentation, and data references. The next sections, Using AMP PDRD and CRN Cloud Data Together and Using GP2 and CRN Cloud Data Together, provide step-by-step guidance for setting up your own workspace, incorporating data from these programs, and running analyses across your selected datasets.
AMP PDRD Resources
Getting Started Workspaces
- Tier 1 – Clinical Access
- Tier 2 – Clinical and Omics Access
- Tier 2 – Autoantibodies
- Tier 2 – Proteomics
Data Catalogs
- Tier 1
- Tier 2
ASAP CRN Cloud Resources
The ASAP CRN Cloud is a centralized platform designed for storing, analyzing, and sharing CRN 'omics data with the global research community. Its primary goals are to create an ecosystem for the curation and distribution of these datasets, promote reproducible science, and enable researchers to access and perform meta-analyses on harmonized data in one place. At the time of this pilot launch, the platform hosts a substantial collection of data derived from postmortem brain tissue, which can be explored through the data catalog linked below.
Getting Started Workspace
Data Catalogs
Additional Resources
Using AMP PDRD and CRN Cloud Data Together
Before You Begin
In order to run the following example of AMP PDRD and ASAP CRN Cloud co-analysis, you will need to have access to both repositories, with a signed Data Use Agreement. Registration for both programs must be done with the same Google account you are using to log into Verily Workbench. If you do not have access to both repositories, the following links provide instructions for registration:
In addition to having access to both repositories, users must have access to billing configured in Verily workbench. In Verily Workbench, Pods are created that are associated with cloud billing accounts, and when Workspaces and Data Collections are created, they are associated with Pods. This allows any cloud service costs generated during the use of Verily Workbench to be charged to the correct billing account.
ASAP CRN Cloud and AMP PDRD have both made Pods available to users, and many organizations that use Verily Workbench will have billing accounts and Pods set up. If you do not see any Pods available to use in Verily Workbench, contact your administration to determine how to request access. Please note that with the ASAP CRN Cloud and AMP PDRD programs providing pilot users access to the billing pods, we request that each pilot user is careful and cognizant of their intended cloud billing charges.
For more information about how to set up billing with pods in verily workbench, please see the following link:
Create a New Workspace in Verily Workbench
Create your own new blank workspace and title it:
Assign a title and description, and select the pod you wish to associate this workspace with; click Next
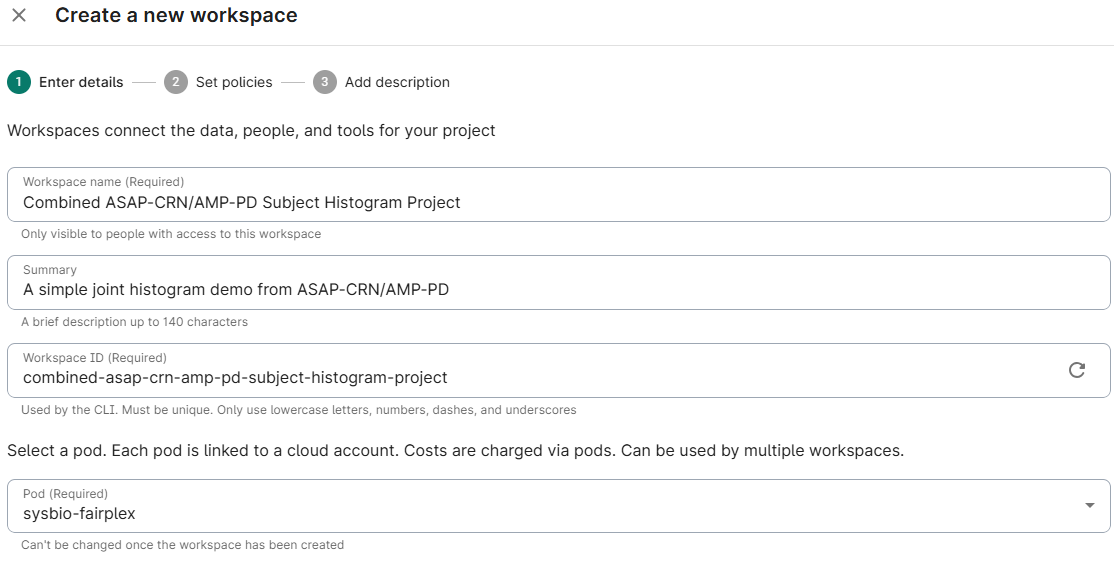
- Choose whether to limit access to groups
- Select the bucket region for your workspace; click Next
- Optionally add a description; click Create Workspace
Add ASAP-CRN Resources to Workspace
Click the + Data from Catalog button
Select ASAP CRN Harmonized Data Collection from the list of available Collections; click Next
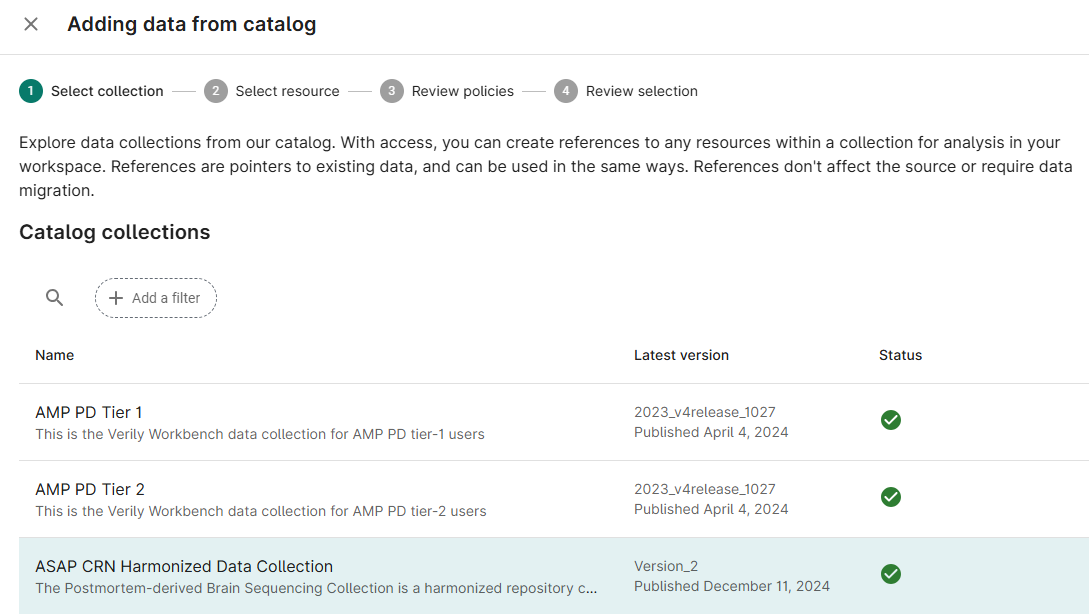
Select checkboxes for the Resource(s) needed for your analysis; click Next
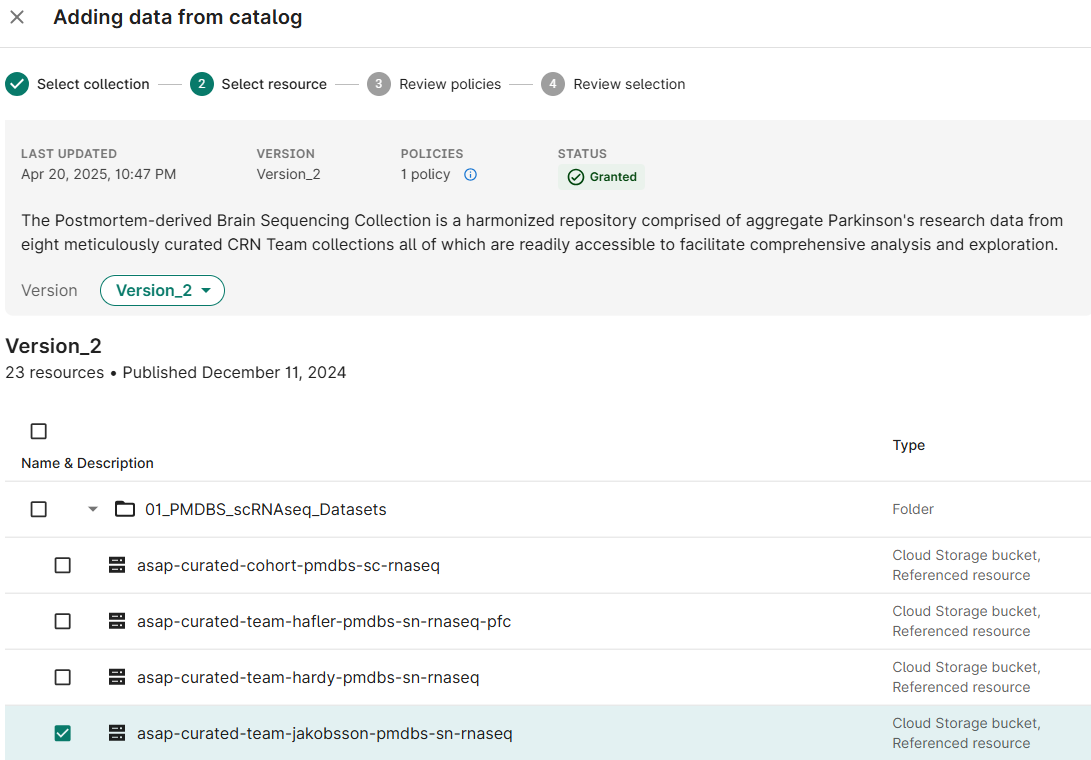
- Accept the policy acknowledgement statement; click Next
- Choose Add to an existing Folder; click Add to your workspace
Add AMP-PD Resources to Workspace
Click the + Data from Catalog button
- Select Pilot - AMP PD Tier 1 from the list of available Collections; click Next
Select checkboxes for the Resource(s) needed for your analysis (gs_path_amp_pd_tier1_release); click Next
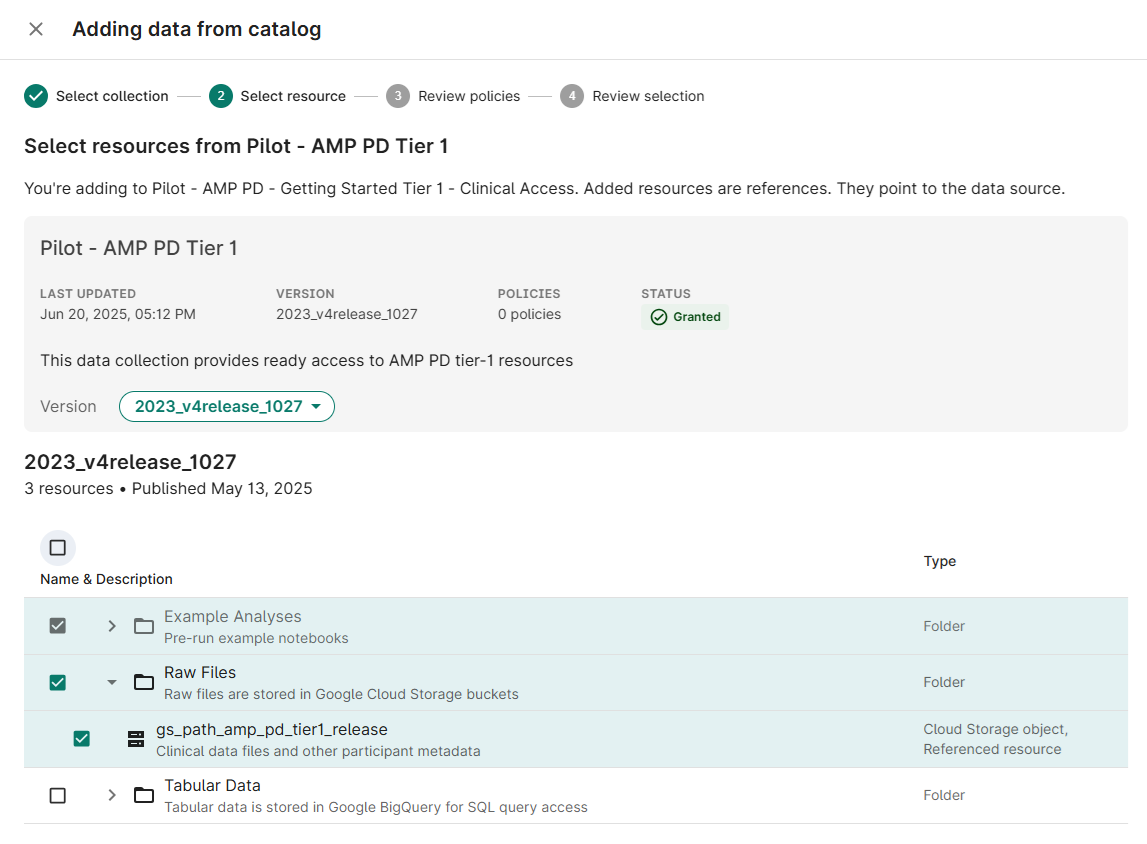
- (If prompted) Accept the policy acknowledgement statement; click Next
- Choose Add to an existing Folder; click Add to your workspace
Select Files for Analysis
From AMP-PD Resource
Select the AMP-PD resource and choose Browse from the info panel
- Navigate to the clinical folder; click Demographics
- Click Add as reference from the info panel
- Accept defaults; click Add to resources
- (Optionally) Choose additional resources
- Click Close to return to the Resources page
From ASAP-CRN Resource
Select the ASAP-CRN resource and choose Browse from the info panel
- Navigate to the metadata → release folder of any dataset; click SUBJECT
- Click Add as reference from the info panel
- Accept defaults; click Add to resources
- (Optionally) Choose additional resources
- Click Close to return to the Resources page
Create an Output Bucket
From the Resources tab, click + New Resource
Select New Cloud Storage Bucket
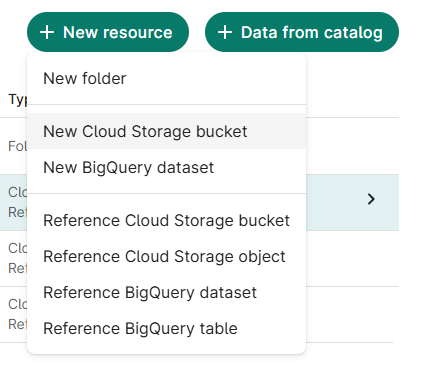
- Assign resource ID; click Create bucket
Create a Virtual Machine to Run Analysis
From the Apps tab, click New app instance, select JupyterLab; click Next
- Choose Default configuration; click Next
- (Optionally) Reduce Machine Type to n1-standard-1
(Optionally) Reduce Data disk size to 100 GB
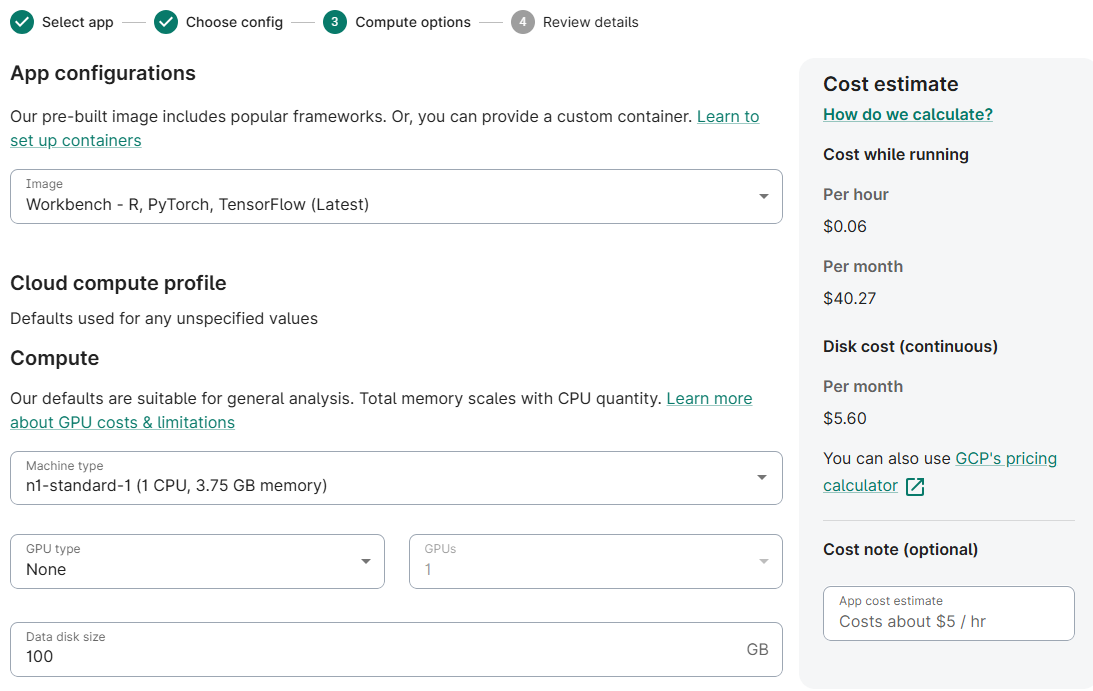
- Accept defaults; click Next
- Review and confirm the app summary; click Create App
- (Note) This step may take several minutes
Create a Notebook for Combined Analysis
From the Apps tab, Launch the newly created virtual machine
- In the JupyterLab Interface that opens, click the Folder Icon in the left menu bar
- In the file browser panel, select the Output Bucket you created earlier.
- In the bucket contents section, right click and select New Notebook
- Select Python 3 (local) kernel type; click Next
Configure Notebook to Analyse Selected Resources
Import Required Libraries
# Use the os package to interact with the environment
import os
# Bring in Pandas for Dataframe functionality
import pandas as pd
# Use StringIO for working with file contents
from io import StringIO
# Enable IPython to display matplotlib graphs.
import matplotlib.pyplot as plt
%matplotlib inline
Create Utility Routines
# Utility routines for reading files from Google Cloud Storage
def gcs_read_file(path):
"""Return the contents of a file in GCS"""
contents = !gsutil -u {BILLING_PROJECT_ID} cat {path}
return '\n'.join(contents)
Read GCS Locations from Workspace Resources
# Get the GCP billing project ID from workbench environment variables
OUTPUT = !wb utility execute env | grep "GOOGLE_CLOUD_PROJECT" | cut -d"=" -f2
BILLING_PROJECT_ID = OUTPUT[0]
# Get the AMP PD Demographics file location from workbench environment variables
OUTPUT = !wb resource resolve --name=Demographics-csv
GS_AMP_PD_DATA_PATH = OUTPUT[0]
# Get the ASAP CRN Subject file location from workbench environment variables
OUTPUT = !wb resource resolve --name=SUBJECT-csv
GS_ASAP_CRN_DATA_PATH = OUTPUT[0]
print(f'BILLING_PROJECT_ID = {BILLING_PROJECT_ID}')
print(f'GS_AMP_PD_DATA_PATH = {GS_AMP_PD_DATA_PATH}')
print(f'GS_ASAP_CRN_DATA_PATH = {GS_ASAP_CRN_DATA_PATH}')

Read AMP PD Demographics Data
# Read CSV file from GCS into dataframe
amp_pd_demographics_df = pd.read_csv(StringIO(gcs_read_file(GS_AMP_PD_DATA_PATH)), engine='python', index_col=False)
amp_pd_demographics_df.head()
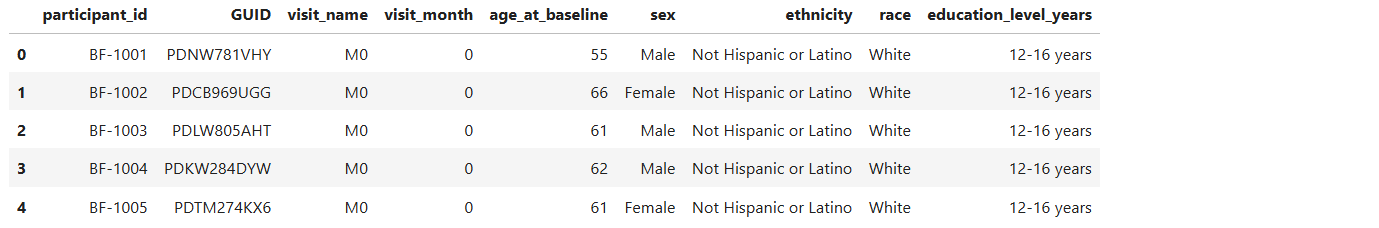
Read ASAP CRN Subject Data
# Read CSV file from GCS into dataframe
asap_crn_subjects_df = pd.read_csv(StringIO(gcs_read_file(GS_ASAP_CRN_DATA_PATH)), engine='python', index_col=[0])
asap_crn_subjects_df.head()
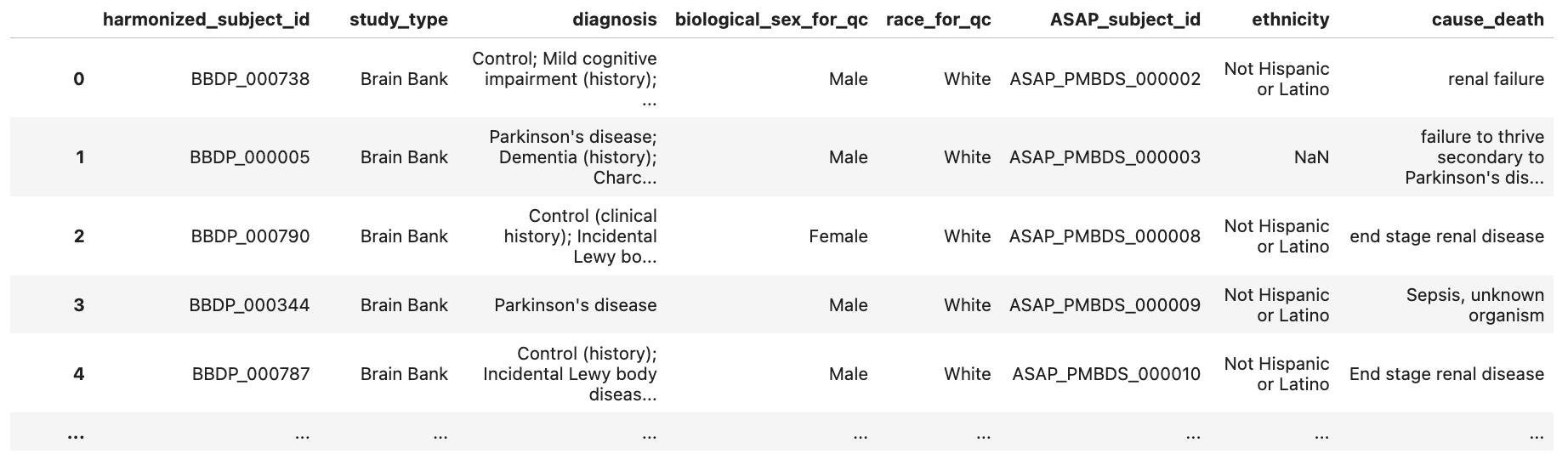
Harmonize and Merge AMP PD and ASAP CRN Data
# rename columns to harmonize data
amp_pd_demographics_df.rename(columns={'participant_id': 'harmonized_subject_id', 'age_at_baseline': 'harmonized_age'}, inplace=True)
asap_crn_subjects_df.rename(columns={'subject_id': 'harmonized_subject_id', 'age_at_collection': 'harmonized_age'}, inplace=True)
# Concatenate dataframes to create new harmonized dataframe
harmonized_df = pd.concat([amp_pd_demographics_df, asap_crn_subjects_df])
harmonized_df
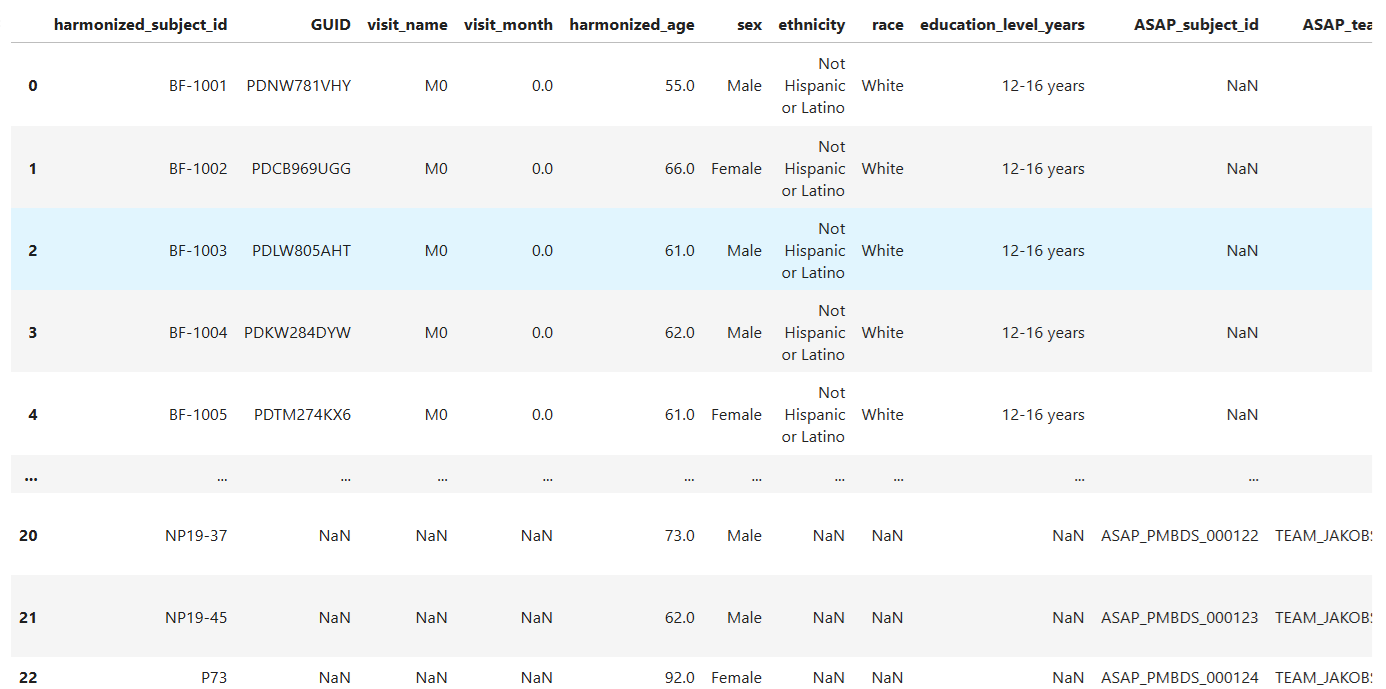
Create Age Distribution Bar Chart
# Configure plot
fig = plt.figure()
fig.set_figheight(6)
fig.set_figwidth(10)
# Create plot
plt.hist(harmonized_df['harmonized_age'].dropna(), bins=20)
plt.xlabel('Age')
plt.ylabel('Number of Participants')
plt.show()
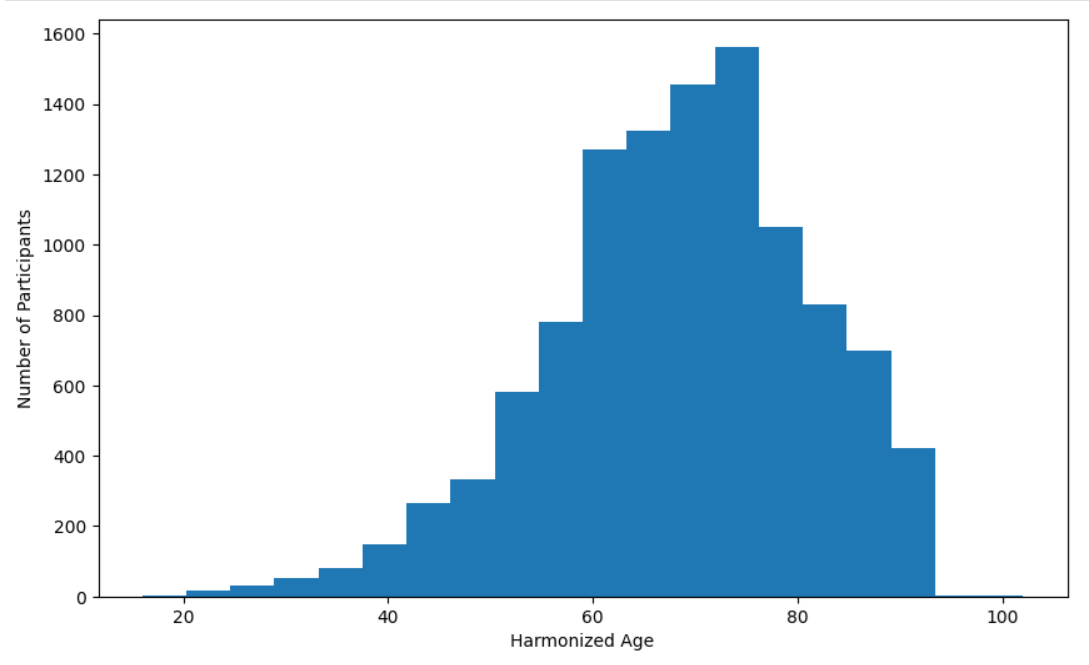
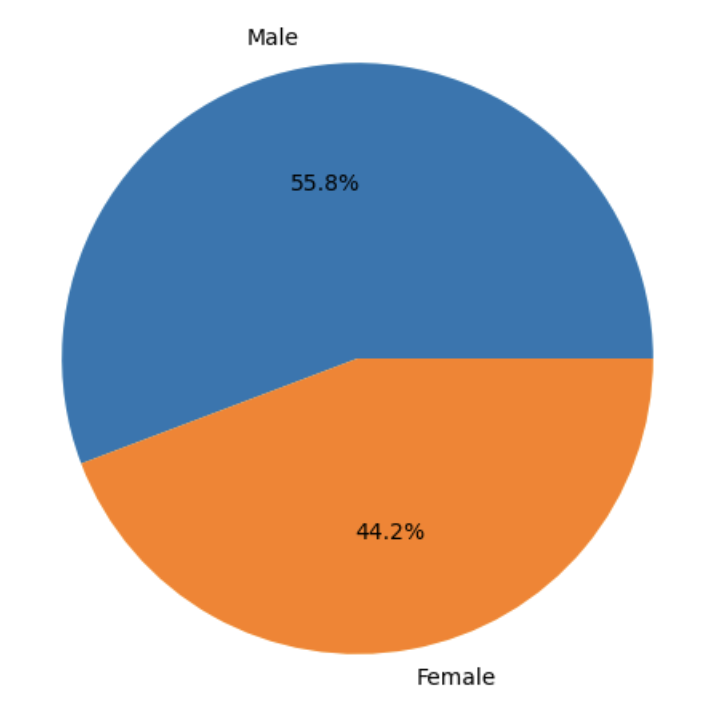
Create Sex Distribution Pie Chart
# Configure plot
counts_by_sex = harmonized_df['sex'].value_counts()
fig = plt.figure()
fig.set_figheight(6)
fig.set_figwidth(10)
# Create plot
plt.pie(counts_by_sex.values, labels=counts_by_sex.index, autopct="%1.1f%%")
plt.figure(figsize=(10, 6))
plt.show()
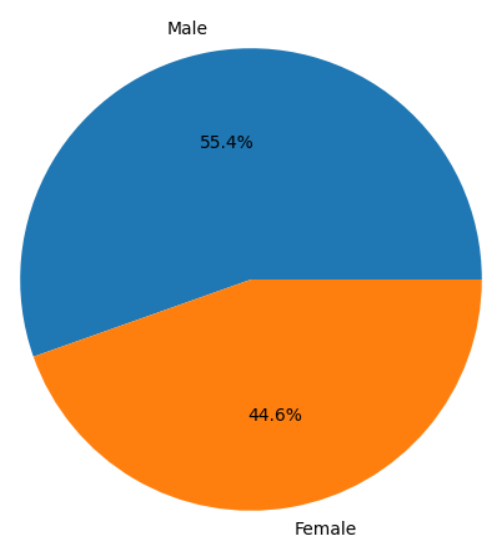
Using GP2 and CRN Cloud Data Together
Before You Begin
In order to run the following example of GP2 and ASAP CRN Cloud co-analysis, you will need to have access to both repositories, with a signed Data Use Agreement. Registration for both programs must be done with the same Google account you are using to log into Verily Workbench. Access to GP2 data is granted through AMP PDRD Registration. If you do not have access to both repositories, the following links provide instructions for registration:
In addition to having access to both repositories, users must have access to billing configured in Verily workbench. In Verily Workbench, Pods are created that are associated with cloud billing accounts, and when Workspaces and Data Collections are created, they are associated with Pods. This allows any cloud service costs generated during the use of Verily Workbench to be charged to the correct billing account.
GP2 has a billing Pod available for users who are part of an approved GP2 project. If you do not see any Pods available to use in Verily Workbench, contact your administration to determine how to request access.
For more information about how to set up billing with pods in verily workbench, please see the following link:
To browse projects currently approved by GP2 and to apply for an approved GP2 project please see the following links:
Create a New Workspace in Verily Workbench
Per GDPR compliance requirements, GP2 data can only be accessed within a perimeter. In order to use GP2 data, your workspace must be created in or added to the perimeter. This can be done by creating a workspace by duplicating the available GP2 Tier 1 or Tier 2 official Getting Started workspaces or by creating a new workspace configured with the required regional and access controls.
To populate GP2 data in a new workspace, the workspace must be created in the europe-west4 (Netherlands) region, and a permission policy must be applied that restricts visibility and access to members of the appropriate GP2 access group (GP2_tier1 or GP2_tier2). Workspaces created outside the europe-west4 region or without the required permission policy will not be eligible to host GP2 data.
From the Getting Started workspaces, select the Duplicate option
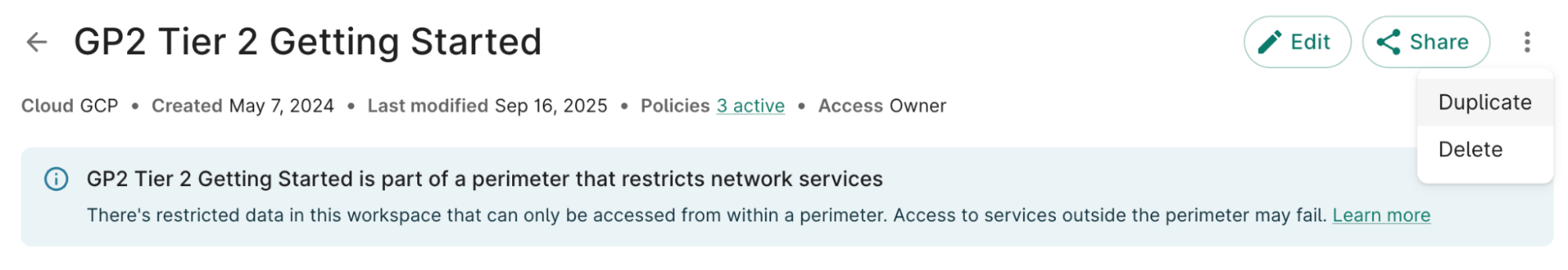
Duplicate the workspace and title it:
Assign a title and description, and select the pod you wish to associate this workspace with; click Next
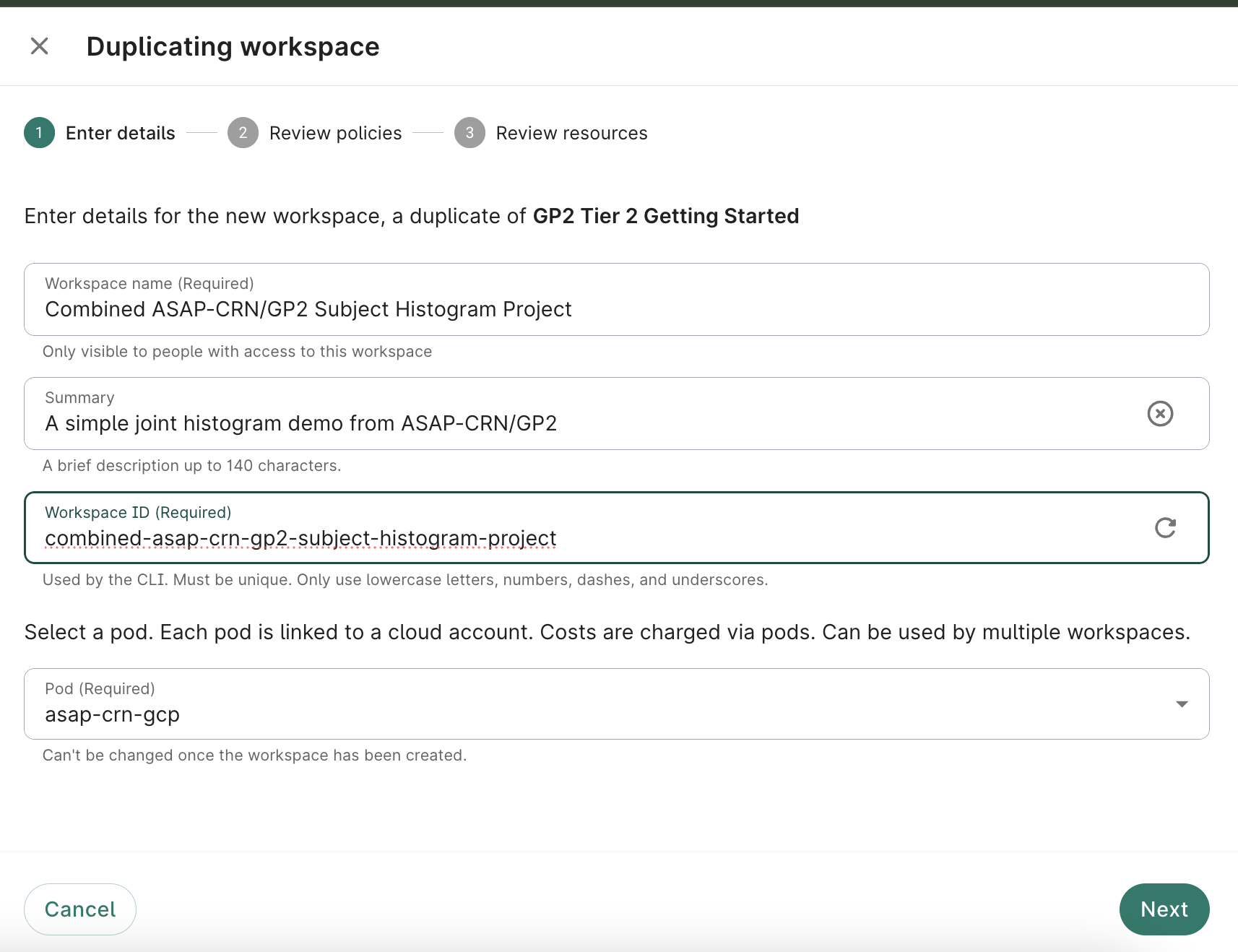
- Ensure access is limited to GP2_Tier1 and GP2_Tier2 groups
Select europe-west4 (Netherlands) as the bucket region; click Next
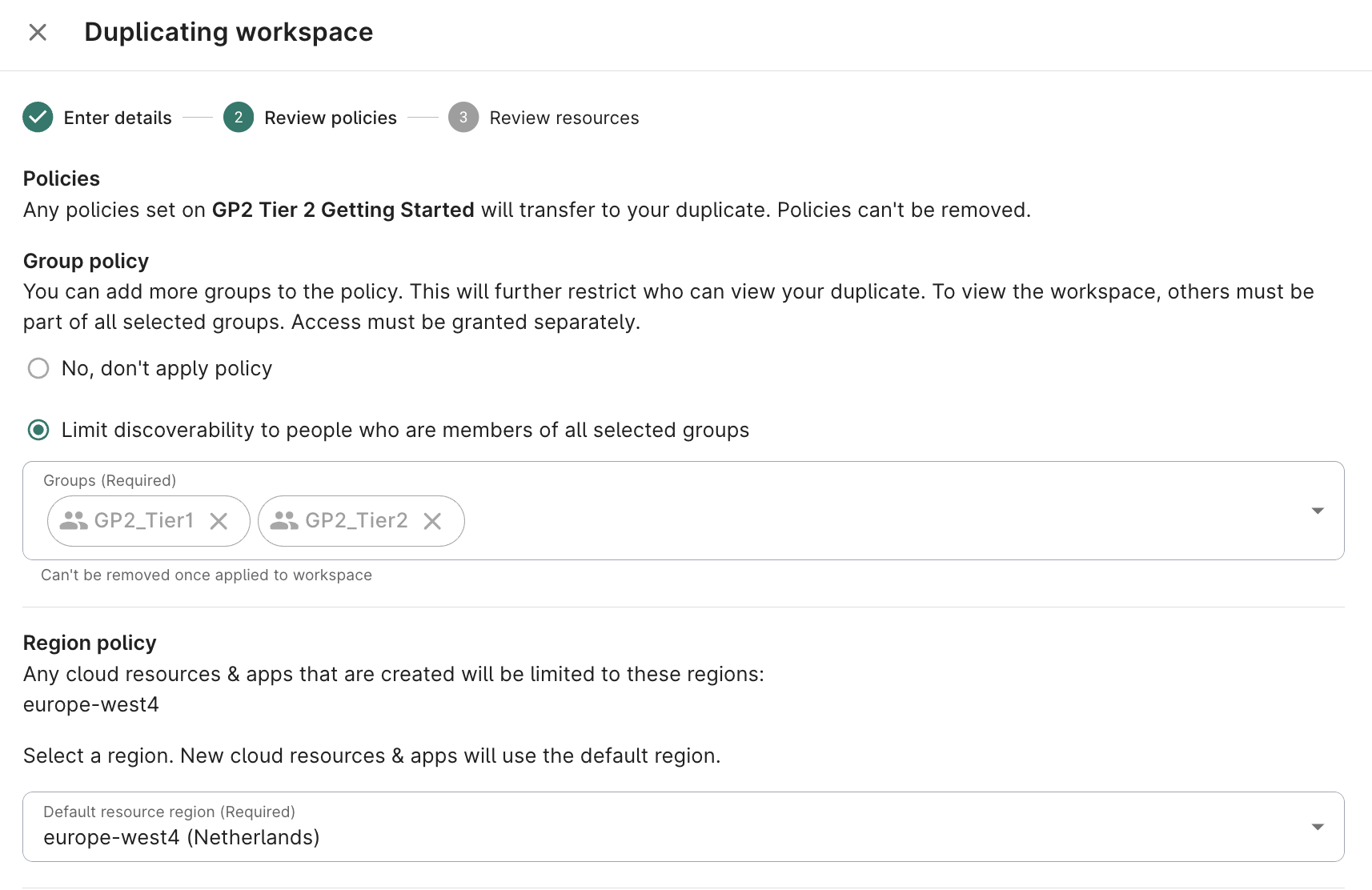
- Optionally add a description; click Create Workspace
Add ASAP-CRN Resources to Workspace
Click the + Data from Catalog button
- Select ASAP CRN Harmonized Data Collection from the list of available Collections; click Next
Select checkboxes for the Resource(s) needed for your analysis; click Next
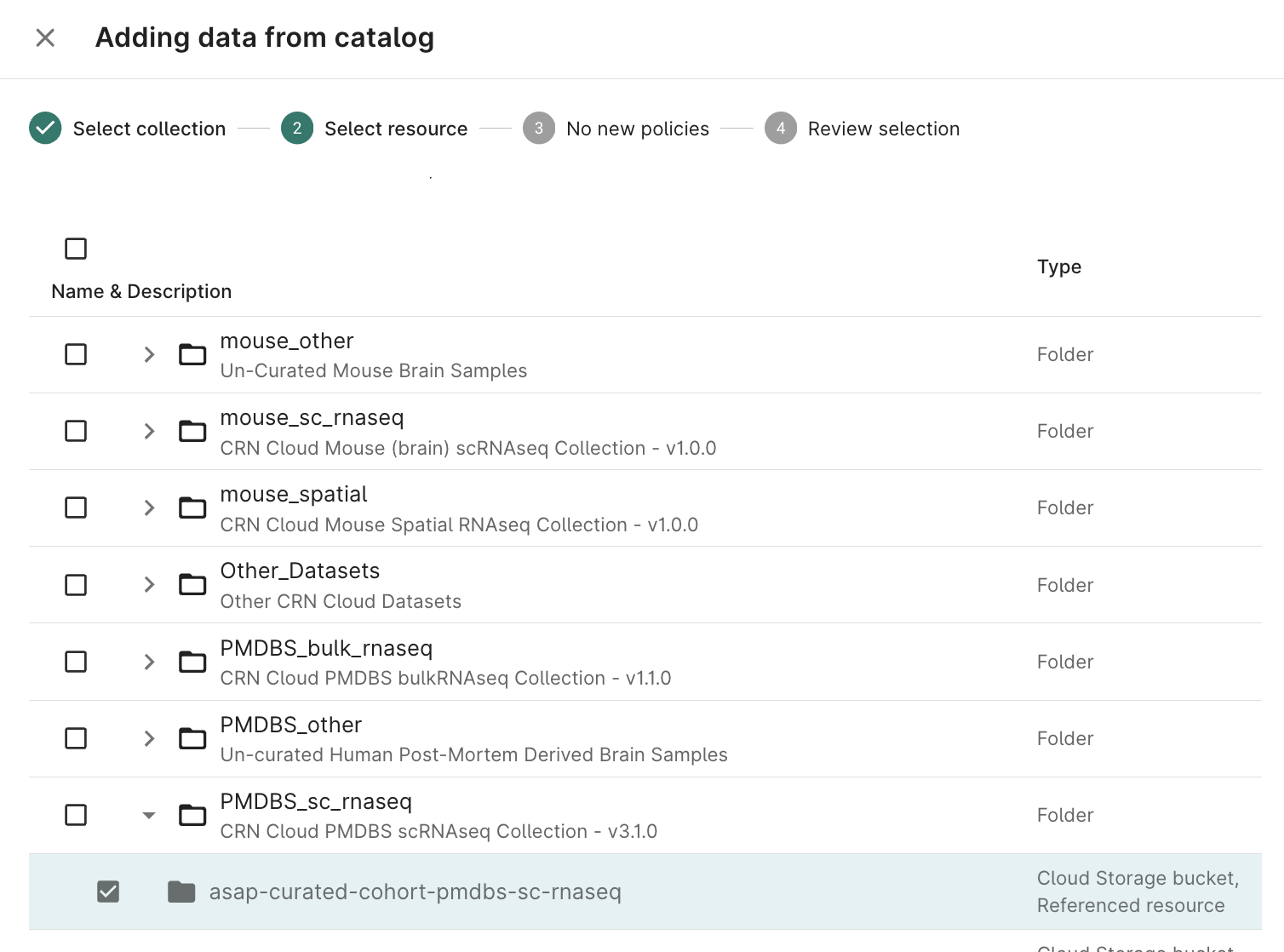
- Accept the policy acknowledgement statement; click Next
- Choose Add to an existing Folder; click Add to your workspace
Select Files for Analysis
From ASAP-CRN Resource
Select the ASAP-CRN resource and choose Browse from the info panel
- Navigate to the metadata → release folder of any dataset; click CLINPATH
- Click Add as reference from the info panel
- Accept defaults; click Add to resources
- (Optionally) Choose additional resources
- Click Close to return to the Resources page
Create an Output Bucket
From the Resources tab, click + New Resource
Select New Cloud Storage Bucket
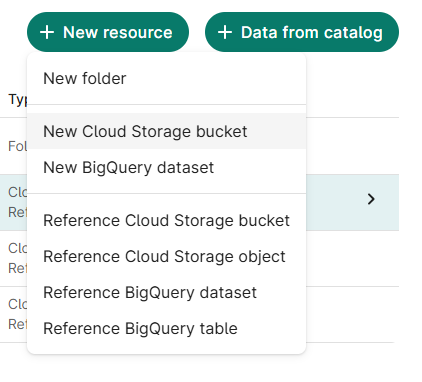
- Assign resource ID; click Create bucket
Create a Virtual Machine to Run Analysis
From the Apps tab, click New app instance, select JupyterLab; click Next
- Choose Default configuration; click Next
- (Optionally) Reduce Machine Type to n1-standard-1
(Optionally) Reduce Data disk size to 100 GB
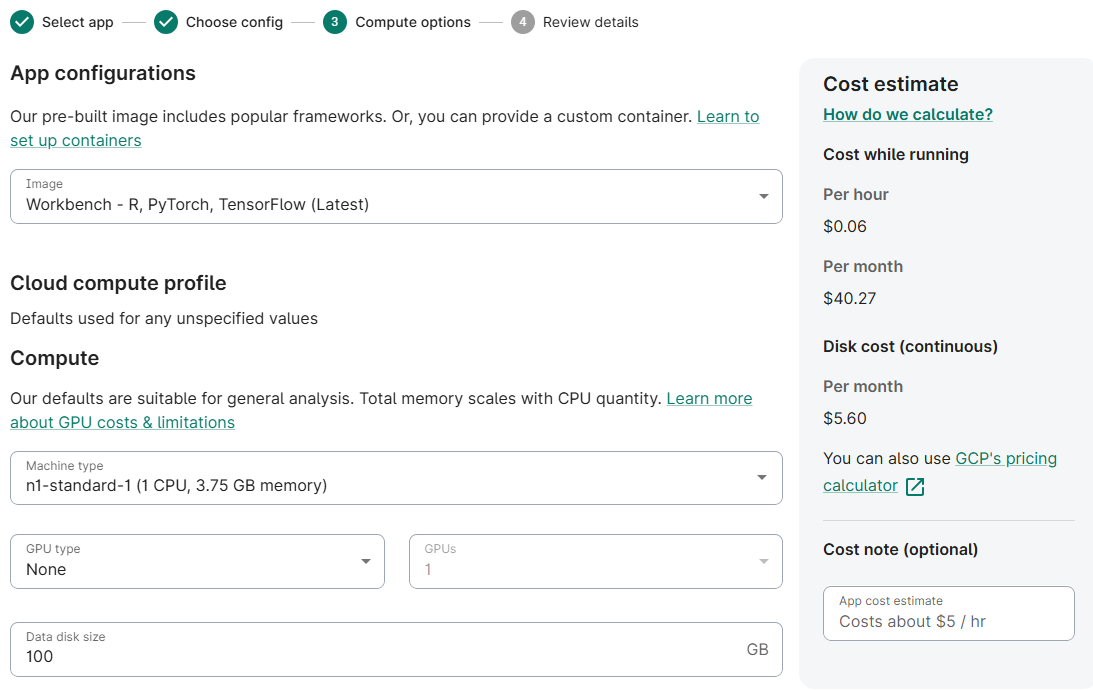
- Accept defaults; click Next
- Review and confirm the app summary; click Create App
- (Note) This step may take several minutes
Create a Notebook for Combined Analysis
From the Apps tab, Launch the newly created virtual machine
- In the JupyterLab Interface that opens, click the Folder Icon in the left menu bar
- In the file browser panel, select the Output Bucket you created earlier.
- In the bucket contents section, right click and select New Notebook
- Select Python 3 (local) kernel type; click Next
Configure Notebook to Analyse Selected Resources
Import Required Libraries
# Use the os package to interact with the environment
import os
# Bring in Pandas for Dataframe functionality
import pandas as pd
# Use StringIO for working with file contents
from io import StringIO
# Enable IPython to display matplotlib graphs.
import matplotlib.pyplot as plt
%matplotlib inline
Create Utility Routines
# Utility routines for reading files from Google Cloud Storage
def gcs_read_file(path):
"""Return the contents of a file in GCS"""
contents = !gsutil -u {BILLING_PROJECT_ID} cat {path}
return '\n'.join(contents)
Read GCS Locations from Workspace Resources
# Get the GCP billing project ID from workbench environment variables
OUTPUT = !wb utility execute env | grep "GOOGLE_CLOUD_PROJECT" | cut -d"=" -f2
BILLING_PROJECT_ID = OUTPUT[0]
# Get the ASAP CRN Subject file location from workbench environment variables
OUTPUT = !wb resource resolve --name=CLINPATH-csv
GS_ASAP_CRN_DATA_PATH = OUTPUT[0]
print(f'BILLING_PROJECT_ID = {BILLING_PROJECT_ID}')
print(f'GS_ASAP_CRN_DATA_PATH = {GS_ASAP_CRN_DATA_PATH}')
Read GP2 Data
# Read CSV file from GCS into dataframe
gp2_data_df = pd.read_csv("workspace/gp2_tier2_eu_release11/clinical_data/master_key_release11_final_vwb.csv", low_memory=False)
gp2_data_df.head()
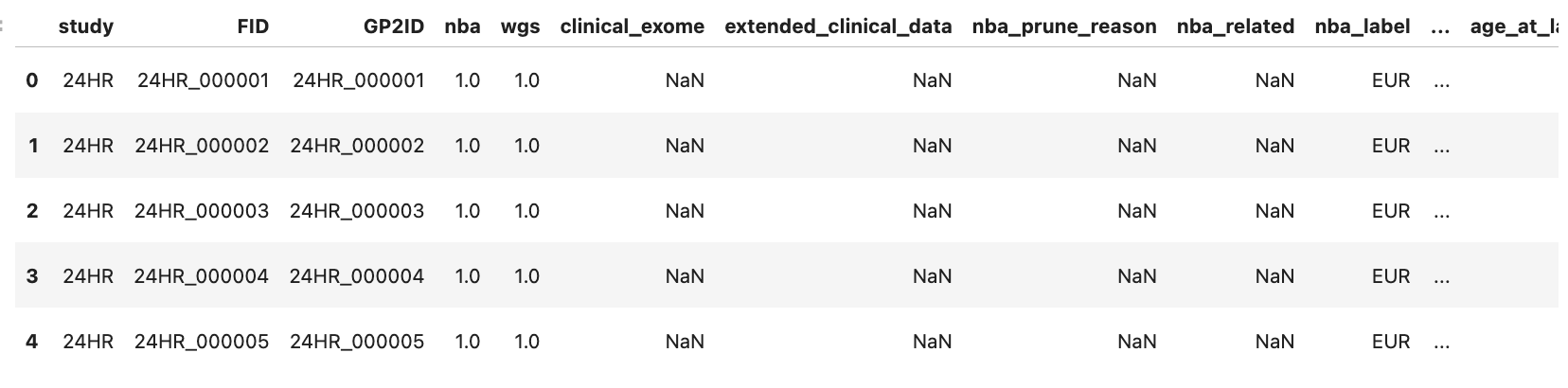
Read ASAP CRN Subject Data
# Read CSV file from GCS into dataframe
asap_crn_subjects_df = pd.read_csv(StringIO(gcs_read_file(GS_ASAP_CRN_DATA_PATH)), engine='python', index_col=[0])
asap_crn_subjects_df.head()
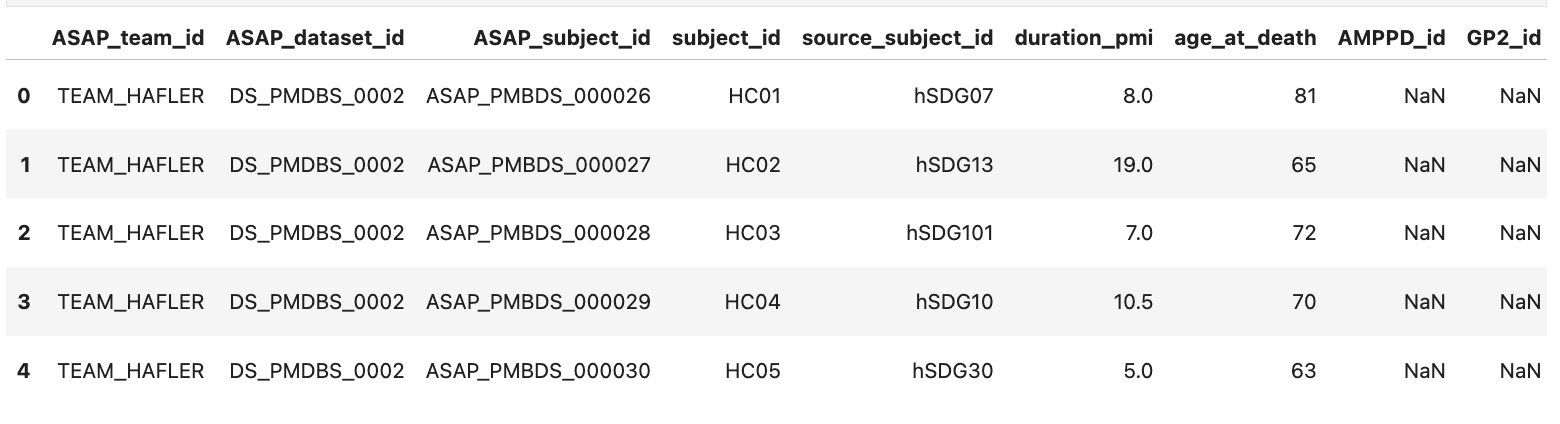
Harmonize and Merge AMP PD and ASAP CRN Data
# rename columns to harmonize data
gp2_data_df.rename(columns={'GP2ID': 'harmonized_subject_id', 'age_at_sample_collection': 'harmonized_age'}, inplace=True)
#removing trailing S1 in GP2 ID for asap cn metadata
asap_crn_clinpath_df['harmonized_subject_id'] = (
asap_crn_clinpath_df['harmonized_subject_id']
.str.replace(r'^(MDGAP.*)_s1$', r'\1', regex=True)
)
asap_crn_clinpath_df.rename(columns={'GP2_id': 'harmonized_subject_id', 'age_at_collection': 'harmonized_age'}, inplace=True)
# Concatenate dataframes to create new harmonized dataframe
harmonized_df = pd.concat([gp2_data_df, asap_crn_clinpath_df])
harmonized_df.head()
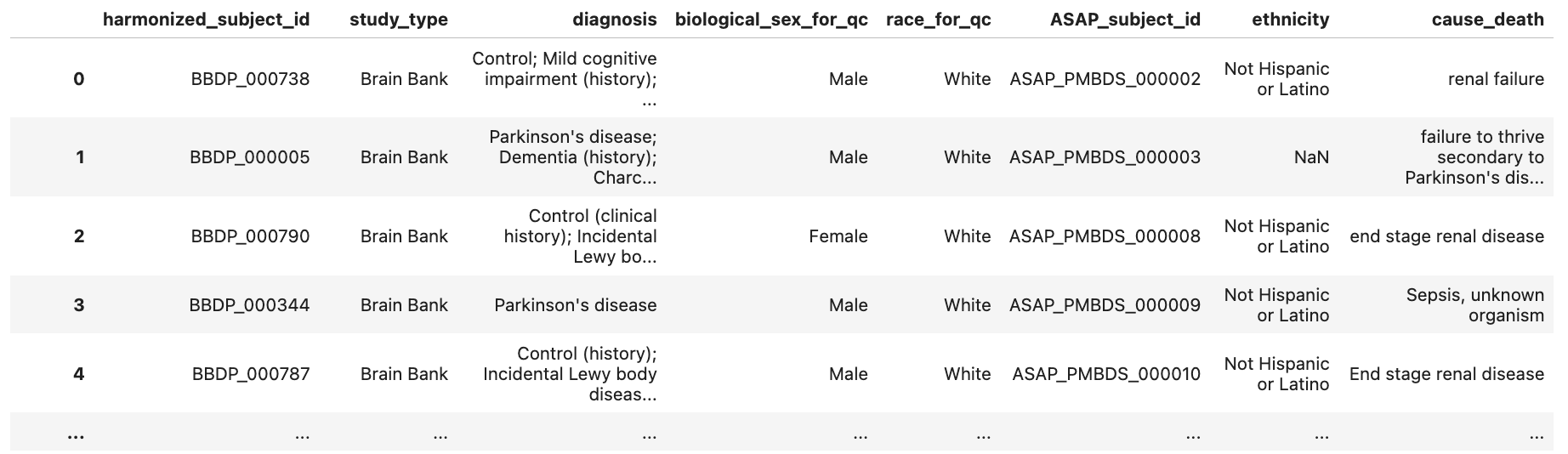
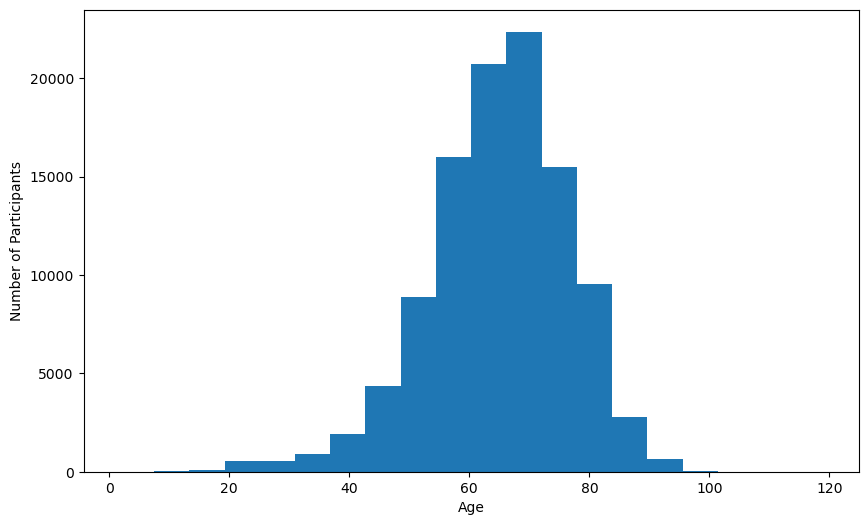
Create Age Distribution Bar Chart
# Configure plot
fig = plt.figure()
fig.set_figheight(6)
fig.set_figwidth(10)
# Create plot
plt.hist(harmonized_df['harmonized_age'].dropna(), bins=20)
plt.xlabel('Age')
plt.ylabel('Number of Participants')
plt.show()
Create Sex Distribution Pie Chart
# Configure plot
counts_by_sex = harmonized_df['biological_sex_for_qc'].value_counts()
fig = plt.figure()
fig.set_figheight(6)
fig.set_figwidth(10)
# Create plot
plt.pie(counts_by_sex.values, labels=counts_by_sex.index, autopct="%1.1f%%")
plt.figure(figsize=(10, 6))
plt.show()